
3D genome organization shapes genetics at all scales. Source: Nature 3D Genome Collection
It’s generally accepted biological knowledge that chromatin is organized in a complex and tightly-regulated manner inside the cellular nucleus. High-throughput chromatin conformation capture (Hi-C), microscopy experiments and computer simulations have added much to our knowledge of chromatin’s three dimensional organization. However, there still isn’t a consensus on the molecular mechanisms actually responsible for such organization!
A major advance in the past few years was the formalization of the loop extrusion hypothesis. Loop extrusion explains many features of experimental data – and I’m going to describe it in detail soon. However, we still don’t have conclusive mechanistic evidence that it is actually happening inside the nucleus. New research is continually adding to the body of evidence for the hypothesis, though, so it looks like loop extrusion is here to stay.
Loop extrusion, explained
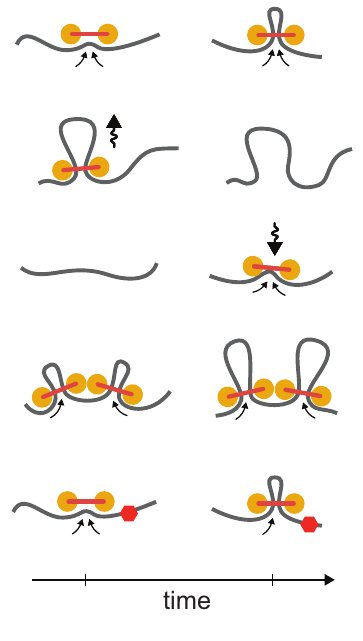
Loop extrusion dynamics. Extruders are shown as linked pairs of yellow circles, chromatin fiber shown in gray. From top to bottom: extrusion, dissociation, association, stalling upon encountering a neighboring extrusion comples, stalling at a blocking element (red hexagon). Taken from Figure 2 of [1]
Loop extrusion considers the action of extruding factors – most likely the proteins cohesin or condensin. Extruding factors are typically thought of as pairs of proteins with a fixed linker between them. They bind DNA and each member of the pair translocates along the polymer, effectively extruding a loop. Unhindered, an extrusion complex will process along the DNA until it dissociates. If an extrusion complex encounters another, both will stall at that position on the DNA. The model also considers blockers – most likely bound CTCF – that can stall extrusion complexes. Blockers are modeled to be directional (they only block extruders moving in one direction) and semi-permeable (they let an extruder pass with a certain probability). For a video explaining the action of loop extrusion, see this post.
Loop extrusion provides a simple unifying principle to explain 3D genome structure. Initially, the best evidence for this hypothesis came from simulation using molecular dynamics – simulating the action of loop extrusion complexes on a polymer constructed to mimic DNA. By placing blocking sites at locations where CTCF was bound in the genome, these simulations closely reconstructed experimental Hi-C data. In addition, loop extrusion can explain what Hi-C contact maps look like after removing CTCF binding sites with CRSIPR genome editing.
There’s a interesting history of the loop extrusion hypothesis – it’s actually not a completely novel idea for explaining genome architecture. A report in 1990 by Arthur Riggs [3] proposed “DNA reeling.” Then, Kim Nasmyth mentioned that an extrusion-like mechanism might play a role in separating sister chromatids and genome organization in 2001 [4]. However, the idea was buried in a 75 page article. Nasmyth even discouraged the idea, “I give this example not so much because it is a serious candidate for the function of condensin (or cohesin for that matter) but rather because it illustrates the notion that condensin or molecules like it could have a very active role in folding and resolving chromatids.” As far as I can tell, loop extrusion went untouched until a publication by Alipour and Marko in 2012 [5], who proposed an extrusion-like mechanism could be used to compact mitotic chromosomes. Then, concurrent publications by the Mirny and Lieberman Aiden groups formalized the idea [1,2]. The timing of these publications was interesting – a preprint of one came out while the other was in review. However, the authors maintain that the ideas were developed independently.
Loop extrusion is a convincing hypothesis, but mechanistic explanation for it actually happening in the genome is lacking. A few recent publications have begun to address this, defining the proteins that are responsible and the parameters that govern their action. For example, Ganji et. al. [6] used single molecule imaging to visualize that condensin from S. cerevisiae could form loops on DNA, at a rate of up to 1500 bp/s. “Here, we provide unambiguous evidence for loop extrusion by directly visualizing the formation and processive extension of DNA loops by yeast condensin in real-time.”
Further biophysical experiments will likely give us more confidence about loop extrusion. In particular, evidence in human cells with human cohesin/condensin is lacking.